Dietary fiber intake and gut microbiota in Chinese patients with type 2 diabetes: secondary data analyses of a randomized controlled trial
-
摘要:
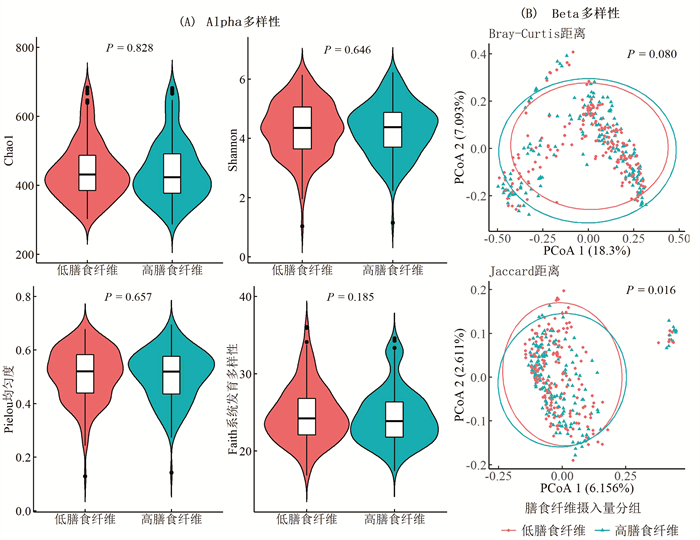
目的 分析2型糖尿病患者膳食纤维摄入量与肠道菌群的关联, 发现相关差异菌群, 为改善患者日常饮食, 调节肠道菌群, 促进健康提供依据。 方法 采用横断面研究设计, 以参与一项健康素养和体力活动干预随机对照试验的356名2型糖尿病患者为研究对象。总热能和膳食纤维摄入量基于3 d 24 h膳食调查数据和中国食物成分表计算。对粪便菌群DNA进行16S rDNA V4高变区测序, 采用Qiime2软件进行生物信息学分析。多变量线性回归模型(multivariate analysis by linear models, MaAsLin)方法和广义线性模型用于获得与膳食纤维摄入量有关的肠道菌群。 结果 研究对象的膳食纤维摄入量处于较低水平, 中位摄入量(四分位间距)仅7.4(5.5, 9.7) g/d。未发现膳食纤维摄入量高低与肠道菌群Alpha多样性有关, 但基于Jaccard距离矩阵(PERMANOVA P=0.016)计算的Beta多样性在两组间差异有统计学意义。低膳食纤维摄入组有较高丰度的梭杆菌属(Fusobacterium)、柯林斯菌属(Collinsella)和普雷沃菌属(Prevotella), 而高膳食纤维摄入组则有较高丰度的黄体链球菌(Streptococcus Luteciae)、脆弱拟杆菌(Fragilis)和双歧杆菌属(Bifidobacterium)。 结论 2型糖尿病患者膳食纤维摄入量与肠道菌群组成有关联。较低膳食纤维摄入量与高丰度的有害菌有关, 而较高摄入量与较高的双歧杆菌属丰度有关, 提示增加膳食纤维摄入可能对患者的肠道菌群和健康有积极作用。 Abstract:Objective To evaluate associations of dietary fiber intake with gut microbiota in Chinese patients with type 2 diabetes (T2DM). Methods A cross-sectional study was conducted among 356 T2DM patients participating in a randomized controlled trial of health literacy and exercise interventions.Total energy and dietary fiber intake were calculated based on the 3-day 24-hour diet recall data and Chinese Food Composition Table.Bacterial 16S rDNA V4 region was sequenced, and Qiime2 was used to conduct bioinformatics analyses.MaAsLin method and generalized linear model were applied to identify differential taxa associated with dietary fiber intake. Results Dietary fiber intake was at a low level in our subjects, with a median intake (IQR) of 7.4(5.5, 9.7) g/day.Beta diversity of fecal microbiota based on the Jaccard distance matrix (PERMANOVA P=0.016) were significantly different between groups with a higher or a lower dietary fiber intake, but no significant association was observed between Alpha diversity and dietary fiber intake.Fusobacterium, Collinsella and Prevotella were more abundant in patients with a lower dietary fiber intake, while Streptococcus Luteciae, Fragilis and Bifidobacterium were enriched in patients taking higher dietary fiber. Conclusions Dietary fiber intake is associated with gut microbiota composition in diabetes patients, with several harmful bacteria abundant in patients taking lower dietary fiber, and Bifidobacterium enriched in those taking higher dietary fiber.Our results suggest potential regulating effect of usual dietary fiber intake on gut microbiota in diabetes patients. -
Key words:
- Type 2 diabetes /
- Dietary fiber /
- Gut microbiota
-
表 1 研究对象基本特征[n(%)]
Table 1. Basic characteristics of subjects [n(%)]
基本特征 所有患者
(N=356)膳食纤维摄入
<7.4 g/d (n=178)膳食纤维摄入
≥7.4 g/d (n=178)P值a P值b 年龄[M(P25, P75), 岁] 65.5(59.0, 71.0) 66.0(60.0, 70.0) 65.0(58.0, 70.0) 0.425 0.192 性别 0.589 0.240 男性 155(43.5) 72(40.4) 83(46.6) 女性 201(56.5) 106(59.6) 95(53.4) 文化程度 0.995 0.676 小学及以下 88(24.7) 48(27.0) 40(22.5) 中学或中专 211(59.3) 100(56.2) 111(62.4) 大专及以上 57(16.0) 30(16.9) 27(15.2) 病程[M(P25, P75), 年] 10(6, 15) 10(7, 15) 10(6, 15) 0.924 0.794 HbA1c[M(P25, P75), %] 7.8(6.9, 9.0) 8.0(7.0, 9.1) 7.8(6.8, 9.0) 0.396 0.376 BMI[M(P25, P75), kg/m2] 24.5(22.9, 26.5) 24.7(23.2, 27.0) 24.4(22.7, 25.9) 0.024 0.029 腰臀比 0.90(0.87, 0.92) 0.90(0.87, 0.92) 0.90(0.88, 0.93) 0.962 0.189 高血压 0.394 0.110 是 244(68.5) 129(72.5) 115(64.6) 否 112(31.5) 49(27.5) 63(35.4) 高血脂 0.127 0.910 是 117(32.9) 59(33.1) 58(32.6) 否 239(67.1) 119(66.9) 120(67.4) 降糖药物c 0.977 0.958 不使用 18(5.1) 8(4.5) 10(5.6) 单用胰岛素 44(12.4) 22(12.4) 22(12.4) 单用口服降糖药 248(69.7) 124(69.7) 124(69.7) 胰岛素+口服降糖药 46(12.9) 24(13.5) 22(12.4) 抗生素c 0.427 0.347 使用 31(8.7) 18(10.1) 13(7.3) 不使用 325(91.3) 160(89.9) 165(92.7) 非甾体抗炎药c 0.004 0.023 使用 69(19.4) 26(14.6) 43(24.2) 不使用 287(80.6) 152(85.4) 135(75.8) 总体力活动[M(P25, P75), MET-h/周] 60(34, 88) 63(33, 90) 59(34, 88) 0.859 0.716 总能量摄入[M(P25, P75), kcal/d] 1 843(1 402, 2 394) 1 675(1 306, 2 346) 1 987(1 537, 2 410) 0.004 0.004 膳食纤维摄入[M(P25, P75), g/d] 7.4(5.5, 9.7) 5.5(4.3, 6.5) 9.7(8.4, 11.6) - - 注:a为简单线性回归P值,膳食纤维摄入量作为连续变量进行分析,基本特征中的分类变量以哑变量形式纳入模型;b为Wilcoxon检验或X2检验的P值; c表示粪便收集时近6个月用药。 表 2 膳食纤维摄入量高低与相关肠道菌群丰度的关联
Table 2. Associations of dietary fiber intake with abundances of related taxa
门 纲 目 科 属/种 丰度(%) β(95% CI)值a P值 Q值 放线菌门 放线菌纲 双歧杆菌目 双歧杆菌科 双歧杆菌属(未知种) 2.630(2.286~3.210) 0.033(0.004~0.062) 0.027 0.399 放线菌门 红蝽菌纲 2.172(1.842~2.528) -0.022(-0.038~-0.007) 0.004 0.085 红椿菌目 2.172(1.842~2.528) -0.022(-0.038~-0.007) 0.004 0.131 红椿菌目科 2.172(1.842~2.528) -0.022(-0.038~-0.007) 0.004 0.253 柯林斯菌属 1.829(1.491~2.307) -0.031(-0.048~-0.015) <0.001 0.030 产气柯林斯菌 1.699(1.342~2.196) -0.027(-0.045~-0.010) 0.002 0.158 拟杆菌门 拟杆菌纲 拟杆菌目 帕拉普雷沃菌科 普雷沃菌属 1.740(1.114~2.064) -0.034(-0.061~-0.007) 0.013 0.309 普雷沃菌科 普雷沃菌属(未知种) 2.236(1.911~2.687) -0.022(-0.042~-0.002) 0.034 0.430 厚壁菌门 4.778(4.645~4.866) -0.006(-0.012~-0.001) 0.032 0.193 芽孢杆菌纲 乳杆菌目 链球菌科 链球菌属(未知种) 0.301(0.000~0.778) -0.029(-0.047~-0.012) 0.001 0.152 黄体链球菌b 0.000(0.000~0.699) 0.026(0.005~0.046) 0.016 0.340 厚壁菌门 梭菌纲 梭菌目 毛螺菌科 多尔氏菌属 2.097(1.767~2.371) -0.017(-0.033~0.000) 0.047 0.504 韦荣氏菌科 巨单胞菌属 2.755(2.580~3.255) -0.030(-0.050~-0.010) 0.003 0.182 梭杆菌门b 2.416(2.244~2.869) -0.029(-0.050~-0.008) 0.007 0.085 梭杆菌纲b 2.416(2.244~2.869) -0.029(-0.050~-0.008) 0.007 0.085 梭杆菌目b 2.416(2.244~2.869) -0.029(-0.050~-0.008) 0.007 0.131 梭杆菌科b 2.411(2.238~2.869) -0.029(-0.050~-0.008) 0.007 0.261 梭杆菌属b 2.411(2.238~2.869) -0.029(-0.050~-0.008) 0.007 0.217 变形菌门 α-变形菌纲 1.638(1.362~2.137) -0.017(-0.034~0.000) 0.045 0.363 柄杆菌目 0.477(0.000~0.954) -0.020(-0.038~-0.002) 0.032 0.233 柄杆菌科 0.477(0.000~0.954) -0.020(-0.038~-0.002) 0.032 0.448 柄杆菌属 0.000(0.000~0.602) -0.015(-0.028~-0.002) 0.028 0.365 变形菌门 α-变形菌纲 鞘脂单胞菌目 1.000(0.602~1.439) -0.024(-0.043~-0.004) 0.016 0.194 鞘脂单胞菌科 1.000(0.602~1.439) -0.023(-0.042~-0.004) 0.020 0.448 鞘氨醇单胞菌属 0.954(0.602~1.406) -0.023(-0.042~-0.004) 0.020 0.343 鞘氨醇单胞菌属(未知种) 0.845(0.477~1.301) -0.021(-0.040~-0.002) 0.028 0.399 变形菌门 δ-变形菌纲 脱硫弧菌目 脱硫弧菌科 脱硫弧菌属 1.833(1.580~2.467) -0.031(-0.053~-0.009) 0.006 0.216 D168脱硫弧菌 1.060(0.778~1.322) -0.031(-0.052~-0.009) 0.005 0.222 变形菌门 γ-变形菌纲 假单胞菌目 2.196(1.911~2.511) -0.019(-0.035~-0.003) 0.021 0.194 假单胞菌科 2.158(1.892~2.496) -0.018(-0.034~-0.002) 0.027 0.448 注:a为广义线性模型调整年龄、性别、病程、BMI、近6个月降糖药、抗生素、非甾体抗炎药使用、总体力活动、总能量摄入后的β值,表示每增加1g/d膳食纤维摄入,肠道菌群绝对丰度(log10转换)的变化,表中所列为中位数(IQR);b表示与MsAsLin方法发现的差异菌群一致;Q值为FDR矫正P值;展示了P<0.05的全部结果。 -
[1] Lynch SV, Pedersen O. The human intestinal microbiome in health and disease[J]. N Engl J Med, 2016, 375(24): 2369-2379. DOI: 10.1056/NEJMra1600266. [2] Ma Q, Li Y, Li P, et al. Research progress in the relationship between type 2 diabetes mellitus and intestinal flora[J]. Biomed Pharmacother, 2019, 117: 109138. DOI: 10.1016/j.biopha.2019.109138. [3] Makki K, Deehan EC, Walter J, et al. The impact of dietary fiber on gut microbiota in host health and disease[J]. Cell Host Microbe, 2018, 23(6): 705-715. DOI: 10.1016/j.chom.2018.05.012. [4] Ojo O, Feng QQ, Ojo OO, et al. The role of dietary fibre in modulating gut microbiota dysbiosis in patients with type 2 diabetes: a systematic review and meta-analysis of randomised controlled trials[J]. Nutrients, 2020, 12(11): 3239. DOI: 10.3390/nu12113239. [5] Ojo O, Ojo OO, Zand N, et al. The effect of dietary fibre on gut microbiota, lipid profile, and inflammatory markers in patients with type 2 diabetes: a systematic review and meta-analysis of randomised controlled trials[J]. Nutrients, 2021, 13(6): 1805. DOI: 10.3390/nu13061805. [6] Zhao L, Zhang F, Ding X, et al. Gut bacteria selectively promoted by dietary fibers alleviate type 2 diabetes[J]. Science, 2018, 359(6380): 1151-1156. DOI: 10.1126/science.aao5774. [7] Wang L, Fang H, Xia Q, et al. Health literacy and exercise-focused interventions on clinical measurements in Chinese diabetes patients: a cluster randomized controlled trial[J]. E Clinical Medicine, 2019, 17: 100211. DOI: 10.1016/j.eclinm.2019.11.004. [8] Ainsworth BE, Haskell WL, Whitt MC, et al. Compendium of physical activities: an update of activity codes and MET intensities[J]. Med Sci Sports Exerc, 2000, 32(9 Suppl): S498-S504. DOI: 10.1097/00005768-200009001-00009. [9] Bolyen E, Rideout JR, Dillon MR, et al. Reproducible, interactive, scalable and extensible microbiome data science using QⅡME 2[J]. Nat Biotechnol, 2019, 37(8): 852-857. DOI: 10.1038/s41587-019-0209-9. [10] Mallick H, Rahnavard A, Mciver LJ, et al. Multivariable association discovery in population-scale meta-omics studies[J]. PLoS Comput Biol, 2021, 17(11): e1009442. DOI: 10.1371/journal.pcbi.1009442. [11] Koh A, De Vadder F, Kovatcheva-Datchary P, et al. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites[J]. Cell, 2016, 165(6): 1332-1345. DOI: 10.1016/j.cell.2016.05.041. [12] Martin-Gallausiaux C, Marinelli L, Blottière HM, et al. SCFA: mechanisms and functional importance in the gut[J]. Proc Nutr Soc, 2021, 80(1): 37-49. DOI: 10.1017/s0029665120006916. [13] Morrison DJ, Preston T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism[J]. Gut Microbes, 2016, 7(3): 189-200. DOI: 10.1080/19490976.2015.1134082. [14] Deehan EC, Duar RM, Armet AM, et al. Modulation of the gastrointestinal microbiome with nondigestible fermentable carbohydrates to improve human health[J]. Microbiol Spectr, 2017, 5(5): 1-24. DOI: 10.1128/microbiolspec.BAD-0019-2017. [15] 张继国, 王惠君, 王志宏, 等. 中国15省(区、直辖市)成年居民膳食纤维摄入状况[J]. 中国食物与营养, 2018, 24(10): 10-12. https://www.cnki.com.cn/Article/CJFDTOTAL-ZGWY201810002.htmZhang JG, Wang HJ, Wang ZH, et al. Dietary fiber intake status among Chinese adults in 15 provinces in 2015[J]. Food Nutri China, 2018, 24(10): 10-12. https://www.cnki.com.cn/Article/CJFDTOTAL-ZGWY201810002.htm [16] King DE, Mainous AG, Lambourne CA. Trends in dietary fiber intake in the United States, 1999-2008[J]. J Acad Nutr Diet, 2012, 112(5): 642-648. DOI: 10.1016/j.jand.2012.01.019. [17] Segata N. Gut microbiome: westernization and the disappearance of intestinal diversity[J]. Curr Biol, 2015, 25(14): R611-R613. DOI: 10.1016/j.cub.2015.05.040. [18] Cummings JH, Macfarlane GT. The control and consequences of bacterial fermentation in the human colon[J]. J Appl Bacteriol, 1991, 70(6): 443-459. DOI: 10.1111/j.1365-2672.1991.tb02739.x. [19] Windey K, De Preter V, Verbeke K. Relevance of protein fermentation to gut health[J]. Mol Nutr Food Res, 2012, 56(1): 184-196. DOI: 10.1002/mnfr.201100542. [20] Kostic AD, Gevers D, Pedamallu CS, et al. Genomic analysis identifies association of fusobacterium with colorectal carcinoma[J]. Genome Res, 2012, 22(2): 292-298. DOI: 10.1101/gr.126573.111. [21] Gomez-Arango LF, Barrett HL, Wilkinson SA, et al. Low dietary fiber intake increases collinsella abundance in the gut microbiota of overweight and obese pregnant women[J]. Gut Microbes, 2018, 9(3): 189-201. DOI: 10.1080/19490976.2017.1406584. [22] Barrett HL, Gomez-Arango LF, Wilkinson SA, et al. A vegetarian diet is a major determinant of gut microbiota composition in early pregnancy[J]. Nutrients, 2018, 10(7): 890. DOI: 10.3390/nu10070890. [23] Crusell MKW, Hansen TH, Nielsen T, et al. Gestational diabetes is associated with change in the gut microbiota composition in third trimester of pregnancy and postpartum[J]. Microbiome, 2018, 6(1): 89. DOI: 10.1186/s40168-018-0472-x. [24] Wu GD, Chen J, Hoffmann C, et al. Linking long-term dietary patterns with gut microbial enterotypes[J]. Science, 2011, 334(6052): 105-108. DOI: 10.1126/science.1208344. [25] Larsen JM. The immune response to prevotella bacteria in chronic inflammatory disease[J]. Immunology, 2017, 151(4): 363-374. DOI: 10.1111/imm.12760. [26] Medina-Vera I, Sanchez-Tapia M, Noriega-López L, et al. A dietary intervention with functional foods reduces metabolic endotoxaemia and attenuates biochemical abnormalities by modifying faecal microbiota in people with type 2 diabetes[J]. Diabetes Metab, 2019, 45(2): 122-131. DOI: 10.1016/j.diabet.2018.09.004. [27] Slavin J. Fiber and prebiotics: mechanisms and health benefits[J]. Nutrients, 2013, 5(4): 1417-1435. DOI: 10.3390/nu5041417. [28] Liang X, Li H, Tian G, et al. Dynamic microbe and molecule networks in a mouse model of colitis-associated colorectal cancer[J]. Sci Rep, 2014, 4: 4985. DOI: 10.1038/srep04985. [29] Wong SH, Yu J. Gut microbiota in colorectal cancer: mechanisms of action and clinical applications[J]. Nat Rev Gastroenterol Hepatol, 2019, 16(11): 690-704. DOI: 10.1038/s41575-019-0209-8. -





 下载:
下载: